Revolutionary AI Model Boosts Protein Interaction Predictions by 17%, Aiding Drug Discovery
April 20, 2026
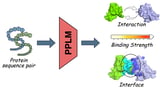
The paired protein language model (PPLM) from the National University of Singapore is trained on over three million protein pairs to learn inter-protein relationships and interaction patterns at scale, enabling predictions of whether proteins interact, their binding strength, and interaction interfaces.
PPLM introduces three specialized tools—PPLM-PPI for predicting protein–protein interactions, PPLM-Affinity for estimating binding strength, and PPLM-Contact for identifying interaction interfaces—with up to about a 17% gain in interaction-prediction accuracy over leading methods across multiple species.
In short, PPLM jointly encodes paired protein sequences rather than analyzing single proteins, advancing beyond traditional single-protein analyses to an interaction-aware framework.
The study, published in Nature Communications on March 10, 2026, marks a shift toward interaction-aware modelling with potential to enable proteome-scale interaction discovery and AI-guided therapeutic design.
This work points to applications in drug target identification and therapeutic development, leveraging proteome-scale interaction discovery and prospects for proteome-wide interaction mapping.
Future work envisions integrating structural and experimental data and extending the approach to more complex systems, such as host–pathogen interactions, to broaden translational impact in drug discovery and disease understanding.
Authors emphasize ongoing work to expand applications to complex biological systems and to translate findings into scalable insights for drug discovery and disease research.
Led by Professor Zhang Yang at the National University of Singapore, the research team developed PPLM, a novel AI model that predicts protein–protein interactions by jointly encoding paired protein sequences.
PPLM’s design contrasts with conventional methods by analyzing paired sequences, enabling more accurate predictions of how proteins interact.
PPLM outperforms traditional sequence-based and structure-based methods, achieving up to 17% higher accuracy and performing strongly across multiple species, including challenging antibody–antigen interactions.
The model captures biologically meaningful interaction patterns that align with real protein interactions, demonstrating robust performance in difficult scenarios like antibody–antigen binding.
Summary based on 2 sources